

Biotempo is a computer science project funded by ANR Blanc (SIMI2) call in 2010. It started in march 2011.
Software (Biotempo Mobyle server) : www
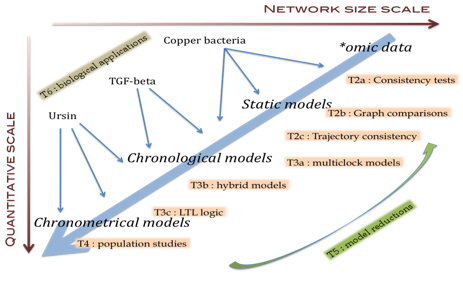
The panel of modeling approaches that are available currently is unfortunately quite poor with respect to the concept of time. Roughly, models can be static (time is not considered, this is mainly the core of integrative biology), chronological (time is discrete, heart of model checking approaches) or chronometric (time is continuous, leading to differential equations, with a strong need of parameters). In these approaches, the representation of time is correlated to the representation of products activity (discrete/discrete or continuous/continuous), which prevents to take into account a large amount of observation and also prevents to switch easily from one model to the other.
Our goal is to combine and complete these three approaches (static, chronological and chronometric) in order to gain in the analysis and interpretation of biological data. To that purpose, we propose to refine the concept of time and disjoin it from the product representation.
Unifying these approaches in a continuous hierarchy is also a strong purpose of this project. We propose to handle that question with two complementary approaches
Three biological questions will be considered to test and validate the new formalisms that we will develop in this project. The three biological questions we wish to apply our development on are a good illustration of the range of data and questions raised by molecular biology.
The goal of this task is to develop both new algorithms (based on graph theory) and constrained-based approaches to address the integration of dynamical-nature data in the process of regulatory networks construction. Constrained-based approaches will be used mainly in model construction and correction with the help of dynamical omic data (mainly transcriptome data), while algorithms on graphs are needed to take into account the (dynamical) information contained in other networks (metabolic networks or genome maps).
The works of this task aim at providing tools for the analysis of real biological systems where time reveals to be a major parameter. Characterization of the time may be either through some ordered sequences of discrete events occurrences or delayed variations of concurrent lasting actions. The common feature for both the discrete and the hybrid approaches stays in the intended algebraic treatment on the obtained timed models : the clock calculus for discrete systems as well as the pi-calculus for hybrid systems are means to apply on the models some operations such as abstraction, equivalence, minimization, emergence of structural properties... These operations are valuable in order to be able to analyze real systems. The result we can obtain are constraints (clock constraints or delay constraints) that have to be checked using qualitative or quantitative temporal logics (e.g. CTL or TCTL) so that the values of a-priori unknown parameters may be specified.
The main objective of this task is to include quantitative reasoning and knowledges in purely quantitative models. For that purpose, it is important to notice that almost every experiments that produce quantitative data corresponds to observations on a cell populations. Quantitative observation thus corresponds to mean and/or variance of certain quantities relative to events over a certain distribution of the population. This task is devoted to the theoretical study of probabilistic models based on weighted Markov chains for modeling quantities like protein concentrations. Such models are constructed by introducing probabilities over discrete chronological models. Quantities are then considered as small accumulations induced by discrete events. The main problems here are to
Computational model reduction, state of the art and beyond Starting from a model with a large number of variables and parameters model reduction allows to obtain a simpler model (less variables and parameters) that can be more easily (faster, precisely) simulated and analyzed. Model reduction in systems biology has some specificity. It is connected with properties of structural incompleteness, parametric incompleteness, multi-scaleness, robustness, combinatorial complexity, stochasticity of models of biochemical networks (signal transduction pathways, metabolic networks). The model reduction techniques in systems biology are usually coarse-graining of networks of biochemical reactions. There exist several classical approaches for model reduction in chemical kinetics (quasi-steady state, quasi-equilibrium approximations, lumping, averaging, finding a limiting rate reaction step). For dealing with large logical and discrete models, existing reduction methods propose suppressing nodes and defining sub-approximating dynamics (Naldi'09). This method belongs to the class of conservative reductions, which preserve essential dynamical properties in terms of attractors and, more importantly, in terms of reachability properties.
This task aims at applying the methods to three real applications.